Immunometabolic characterization of Crohn’s disease phenotypes – Does location matter?
Principal Investigator

Dr. Lea-Maxie Haag
Dr. Malte Lehmann
Scientific interest within the context of the graduate college
Based at the Department of Gastroenterology, Infectious Diseases and Rheumatology at Campus Benjamin Franklin, Charité – Universitätsmedizin Berlin, our main clinical and research focus are inflammatory bowel diseases (IBD). Crohn’s disease (CD), one of the primary forms of IBD, exhibits various clinical phenotypes, including inflammatory (B1), stricturing (B2), and fistulizing (B3) disease phenotypes. We aim at understanding the biological differences between these phenotypes at the level of the mucosal microenvironment. With our large endoscopy unit and outpatient clinic, as well as gastroenterology ward, we have access to biological samples like intestinal biopsies along with the corresponding clinical data.
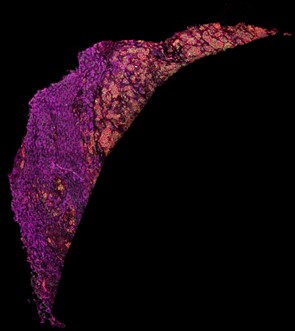
Figure 1. Imaging mass cytometry staining of a liver sample stained with antibodies directed against fatty acid synthase (magenta), voltage-dependent anion-channel 1 (green) and CD44 (red).
Project description
What is the underlying mechanism and biology leading to the different disease locations in Crohn’s disease? This project aims to characterize the immunometabolic landscape of Crohn’s disease depending on disease location. Using an already established 35-marker imaging mass cytometry (IMC) panel, we will analyze mucosal tissue from patients with ileal, colonic, and ileocolonic Crohn’s disease and compare these samples with ulcerative colitis and healthy controls. This approach enables simultaneous analysis of immune, epithelial, and stromal cells at single-cell resolution within their spatial tissue context. In addition to identifying cell populations, it allows investigation of metabolic states, cell-cell interactions, and cellular neighborhoods within the intestinal mucosa.
The project builds on extensive preliminary work from our group, including established IMC workflows, published expertise in spatial single-cell profiling, and an already collected and well-characterized cohort. The required FFPE samples and corresponding clinical data are already available, and all relevant analytical methods are established, making this a highly feasible and well-supported research project.
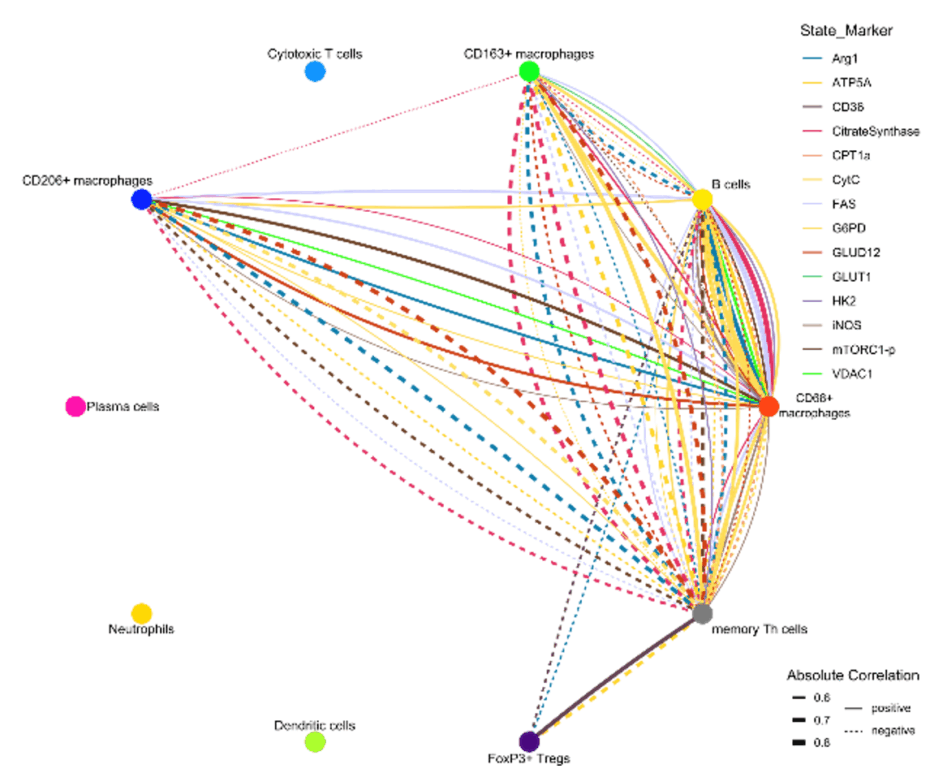
Figure 2. Iconography of correlations of state marker expression to the distance between immune cell clusters. Only correlations with r > 0.5 or < 0.5 and p < 0.05 are shown. The color of the lines indicates the respective state marker, the width of the line the absolute value of correlation. Straight lines indicate positive correlations, doted lines negative correlations.
WP1: Cohort familiarization and sample annotation. Introduction to the clinical cohort and classification of samples from ileal, colonic, and ileocolonic Crohn’s disease, ulcerative colitis, and healthy controls.
WP2: Imaging mass cytometry data generation and analysis. Participation in IMC-based analysis of mucosal biopsies using an established 35-marker panel.
WP3: Spatial and immunometabolic profiling. Analysis of immune cell composition, epithelial and stromal cell states, metabolic marker expression, cell-cell interactions and tissue neighborhoods, differences between disease locations and disease entities.
What the student will do: i) work on a translational IBD research project addressing an important clinical question; ii) analyze high-dimensional spatial single-cell data from human intestinal tissue; iii) compare immunometabolic tissue signatures across Crohn’s disease locations; iv) contribute to interpretation of findings in the context of disease mechanisms and therapy response; v) participate in scientific discussion, presentation of results, and manuscript development.
What the student will learn: i) principles of translational inflammatory bowel disease research in Gastroenterology; ii) application and interpretation of imaging mass cytometry data, iii) spatial single-cell and immunometabolic tissue analysis; iv) R-based analysis workflows for multiplex imaging data; v) scientific writing, presentation, and work in an interdisciplinary clinician-scientist environment.
Supervision and environment. The student will be embedded in an experienced and collaborative research team with close supervision by clinicians and clinician-scientists working at the interface of patient care and experimental research. The group has extensive expertise in inflammatory bowel disease, tissue immunology, and imaging mass cytometry. Weekly meetings, scientific exchange, and structured mentoring will support the student throughout the project.
Expected outcome: i) completion of a MD thesis; ii) strong potential for co-authorship or first authorship, depending on progress and project scope; iii) acquisition of valuable skills for a future clinical-academic career in gastroenterology or immunology.
Target group. This project is ideally suited for a highly motivated medical student seeking a one-year full-time research experience and interested in translational medicine, gastroenterology, immunology, and advanced tissue imaging.
References
- Gajendran M, Loganathan P, Catinella AP, Hashash JG. A comprehensive review and update on Crohn’s disease. Dis Mon. 2018; 64(2): 20-57.
- Atreya R, Siegmund B. Location is important: differentiation between ileal and colonic Crohn’s disease. Nat Rev Gastroenterol Hepatol. 2021; 18(8): 544-558.
- Lehmann M, Weixler B, Elezkurtaj S, Loddenkemper C; TRR241 IBDome Consortium; Kühl AA, Siegmund B. Spatial single cell profiling using imaging mass cytometry: inflammatory versus penetrating Crohn’s disease. J Crohns Colitis. 2024; 18(8): 1305-1318.
- Lehmann M, Allers K, Heldt C, Meinhardt J, Schmidt F, Rodriguez-Sillke Y, Kunkel D, Schumann M, Böttcher C, Stahl-Hennig C, Elezkurtaj S, Bojarski C, Radbruch H, Corman VM, Schneider T, Loddenkemper C, Moos V, Weidinger C, Kühl AA, Siegmund B. Human small intestinal infection by SARS-CoV-2 is characterized by a mucosal infiltration with activated CD8+ T cells. Mucosal Immunol. 2021; 14(6): 1381-1392.
- Stankey CT*, Bourges C*, Haag LM*, Turner-Stokes T, Piedade AP, Palmer-Jones C, Papa I, Silva Dos Santos M, Zhang Q, Cameron AJ, Legrini A, Zhang T, Wood CS, New FN, Randzavola LO, Speidel L, Brown AC, Hall A, Saffioti F, Parkes EC, Edwards W, Direskeneli H, Grayson PC, Jiang L, Merkel PA, Saruhan-Direskeneli G, Sawalha AH, Tombetti E, Quaglia A, Thorburn D, Knight JC, Rochford AP, Murray CD, Divakar P, Green M, Nye E, MacRae JI, Jamieson NB, Skoglund P, Cader MZ, Wallace C, Thomas DC, Lee JC. A disease-associated gene desert directs macrophage inflammation through ETS2. Nature. 2024 Jun; 630(8016): 447-456. *equal contribution
- Avery EG*, Haag LM*, McParland V, Kedziora SM, Zigra GJ, Valdes DS, Kirchner M, Popp O, Geisberger S, Nonn O, Karlsen TV, N’Diaye G, Yarritu A, Bartolomaeus H, Bartolomaeus TUP, Tagiyeva NA, Wimmer MI, Haase N, Zhang YD, Wilhelm A, Grütz G, Tenstad O, Wilck N, Forslund SK, Klopfleisch R, Kühl AA, Atreya R, Kempa S, Mertins P, Siegmund B, Wiig H, Müller DN. Intestinal interstitial fluid isolation provides novel insight into the human host-microbiome interface. Cardiovasc Res. 2025; 121(5): 803-816. *equal contribution