Epithelial acetyl-CoA metabolism as a regulator of intestinal barrier function in IBD
Prinicipal Investigator

Prof. Dr. Britta Siegmund
Dr. Lea-Maxie Haag
Dr. Malte Lehmann
Scientific interest within the context of the graduate college
Inflammatory bowel disease (IBD) is a chronic, multifactorial disorder characterized by immune dysregulation, microbial imbalance, and impaired epithelial barrier function. Intestinal epithelial cells (IECs) play a central role in maintaining gut homeostasis by integrating signals from the microbiota and the immune system. Emerging evidence from immunometabolism highlights that metabolic reprogramming is a key regulator of inflammatory processes. However, IEC-intrinsic metabolic alterations in IBD remain poorly understood.
Within our preliminary work, integrative analyses of human IBD datasets and tissue samples revealed profound metabolic dysregulation in IECs, with the acetyl-CoA biosynthesis pathway emerging as a critical metabolic hub. In particular, reduced expression of key enzymes such as ACLY and ACSS2 suggests that impaired epithelial metabolism may compromise barrier integrity and promote inflammation. Together, these findings support the concept that epithelial metabolic rewiring is a central driver of IBD pathogenesis and a promising target for therapeutic intervention. Our research group at the Department of Gastroenterology, Infectious Diseases and Rheumatology at Charité – Universitätsmedizin Berlin (Campus Benjamin Franklin) and the Max Delbrück Center (MDC) conducts translational research in IBD, leveraging access to a large, deeply phenotyped patient cohort and comprehensive biobanking infrastructure. The project is embedded within the Collaborative Research Centre TRR241, which integrates clinical, experimental, and computational approaches within the IBDome framework to systematically dissect mechanisms of intestinal inflammation. Within this network, we contribute the human translational component, including access to patient samples and multi-omic analyses, while a partner laboratory in Erlangen performs complementary mechanistic studies in murine models. This integrated, cross-site approach enables the direct linking of human disease phenotypes with experimentally defined mechanisms.
Project description
This project aims to define how epithelial metabolic rewiring contributes to intestinal inflammation in human inflammatory bowel disease (IBD), with a particular focus on the acetyl-CoA biosynthesis pathway. Building on our preliminary findings identifying ACLY and ACSS2 as key metabolic regulators in IECs, we will investigate how altered epithelial metabolism affects barrier integrity and mucosal homeostasis in patients. Using biopsies from well-characterized IBD cohorts, we will perform metabolomic profiling of primary IECs and apply stable isotope tracing to reconstruct metabolic pathway activity. Patient-derived intestinal organoids will be used to distinguish epithelial-intrinsic changes from microenvironment-driven effects and to assess responses to inflammatory and microbial stimuli. In parallel, spatial proteomics and interstitial fluid analysis will provide insight into the local signaling environment shaping epithelial metabolism. By integrating these approaches, the project aims to identify key metabolic pathways underlying epithelial dysfunction in IBD and reveal potential targets for therapeutic intervention.
Embedded within the Collaborative Research Centre TRR241, access to well-characterized patient cohorts, high-quality clinical data, and biological samples is assured. All experimental protocols are established, and data acquisition is ongoing. The existing research network, dedicated study teams, and continuous patient recruitment provide a strong infrastructure and ensure close supervision and reliable project execution.
WP1: Characterization of acetyl-CoA pathway components in human IBD. We will analyze ACLY and ACSS2 expression, localization, and phosphorylation in intestinal biopsies from IBD patients across disease states. Using immunostaining, Western blotting, and integration of IBDome and scRNA-seq datasets, we will correlate epithelial acetyl-CoA pathway activity with disease type, inflammation grade, and therapeutic response.
WP2: Functional analysis of epithelial acetyl-CoA metabolism. Patient-derived intestinal organoids will be used to model epithelial-intrinsic metabolic alterations. Pharmacological inhibition of ACLY and ACSS2 will mimic loss-of-function, and metabolic consequences will be assessed by selected metabolic readouts. In parallel, targeted metabolomic profiling will characterize key acetyl-CoA–related pathways in IBD.
WP3: Spatial characterization of epithelial metabolic rewiring in IBD. We will perform spatial proteomics on FFPE samples from the IBDome cohort to map epithelial protein expression in situ, focusing on UC pancolitis stratified by inflammation status. Laser microdissection followed by mass spectrometry will enable analysis of protein abundance and key post-translational modifications.To complement this, isotope-resolved metabolomics will be applied to selected biopsies to assess acetyl-CoA–related metabolic activity. In addition, interstitial fluid analysis will characterize the local signaling environment, including cytokines and metabolites.
Together, these approaches will provide a spatially resolved view of epithelial metabolism and its regulation within the intestinal microenvironment.
What the student will do: i) work on a translational research project investigating epithelial metabolism in human IBD; ii) analyze human intestinal biopsies, primary epithelial cells, and patient-derived organoids; iii) perform and interpret metabolomic and spatial proteomic analyses; iv) contribute to the characterization of metabolic alterations across disease states; v) participate in data integration, scientific discussion, and manuscript preparation.
What the student will learn: i) principles of translational research in IBD and mucosal immunology; ii) hands-on experience with metabolomics, isotope tracing, and spatial proteomics; iii) analysis and interpretation of multi-omic datasets; iv) scientific writing, presentation skills, and critical data interpretation; v) working in an interdisciplinary clinician-scientist environment.
Supervision and environment. The student will be embedded in an experienced and collaborative research team at Charité and MDC, with close supervision by clinician scientists. The project is part of the CRC241 research network, providing access to a multidisciplinary environment integrating clinical, experimental, and computational expertise. Regular meetings, seminars, and structured mentoring will support the student throughout the project.
Expected outcome. i) completion of a MD thesis; ii) strong potential for co-authorship or first authorship, depending on progress; iii) acquisition of key skills for a future clinical-academic career.
Target group. This project is aimed at highly motivated and academically outstanding medical students seeking a dedicated one-year full-time research experience. Candidates should have a strong interest in translational medicine, gastroenterology, and immunometabolism, as well as a willingness to engage with advanced experimental and analytical approaches. Prior research experience is advantageous but not required. A high level of motivation, reliability, and curiosity is essential.
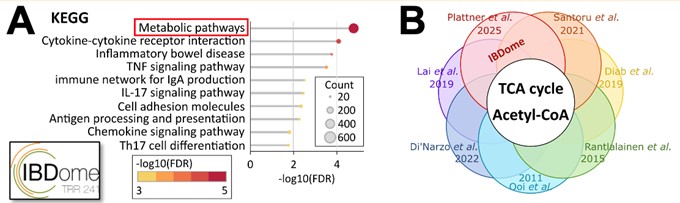
Figure 1. (A) KEGG pathway analysis of differentially expressed genes between IBD patients and controls (IBDome cohort, TRR241). (B) Schematic of overlapping pathways reported in the literature.
References
- Stankey CT*, Bourges C*, Haag LM*, Turner-Stokes T, Piedade AP, Palmer-Jones C, Papa I, Silva Dos Santos M, Zhang Q, Cameron AJ, Legrini A, Zhang T, Wood CS, New FN, Randzavola LO, Speidel L, Brown AC, Hall A, Saffioti F, Parkes EC, Edwards W, Direskeneli H, Grayson PC, Jiang L, Merkel PA, Saruhan-Direskeneli G, Sawalha AH, Tombetti E, Quaglia A, Thorburn D, Knight JC, Rochford AP, Murray CD, Divakar P, Green M, Nye E, MacRae JI, Jamieson NB, Skoglund P, Cader MZ, Wallace C, Thomas DC, Lee JC. A disease-associated gene desert directs macrophage inflammation through ETS2. Nature. 2024; 630(8016): 447-456. *equal contribution
- Avery EG*, Haag LM*, McParland V, Kedziora SM, Zigra GJ, Valdes DS, Kirchner M, Popp O, Geisberger S, Nonn O, Karlsen TV, N’Diaye G, Yarritu A, Bartolomaeus H, Bartolomaeus TUP, Tagiyeva NA, Wimmer MI, Haase N, Zhang YD, Wilhelm A, Grütz G, Tenstad O, Wilck N, Forslund SK, Klopfleisch R, Kühl AA, Atreya R, Kempa S, Mertins P, Siegmund B, Wiig H, Müller DN. Intestinal interstitial fluid isolation provides novel insight into the human host-microbiome interface. Cardiovasc Res. 2025; 121(5): 803-816. *equal contribution
- Santoru ML, Piras C, Murgia F, Leoni VP, Spada M, Murgia A, Liggi S, Lai MA, Usai P, Caboni P, Manzin A, Atzori L. Metabolic alteration in plasma and biopsies from patients with IBD. Inflamm Bowel Dis. 2021; 27(8): 1335-1345.
- Diab J, Hansen T, Goll R, Stenlund H, Jensen E, Moritz T, Florholmen J, Forsdahl G. Mucosal metabolomic profiling and pathway analysis reveal the metabolic signature of ulcerative colitis. Metabolites. 2019; 9(12): 291.
- Smillie CS, Biton M, Ordovas-Montanes J, Sullivan KM, Burgin G, Graham DB, Herbst RH, Rogel N, Slyper M, Waldman J, Sud M, Andrews E, Velonias G, Haber AL, Jagadeesh K, Vickovic S, Yao J, Stevens C, Dionne D, Nguyen LT, Villani AC, Hofree M, Creasey EA, Huang H, Rozenblatt-Rosen O, Garber JJ, Khalili H, Desch AN, Daly MJ, Ananthakrishnan AN, Shalek AK, Xavier RJ, Regev A. Intra- and inter-cellular rewiring of the human colon during ulcerative colitis. Cell. 2019; 178(3): 714-730.e22.