Characterization of the peripheral immune compartment during fasting in healthy volunteers and MS patients
Scientific interest within the context of the graduate college:
Diet is an important factor for a healthy life. For the most part of human history, the next meal was not a given. Hence, there was a strong selection pressure for adaptation to periods of no or low food consumption during our evolution. Consequently, today’s excessive calorie intake, as it is typical for diets in the western world, results in increasing occurrence of systemic inflammation and widespread diseases. In contrast, calorie restriction has been shown to improve numerous chronic diseases and to prolong the healthy lifespan. In my group, we are re-thinking health in the context of evolutionary adaptation to low food energy intake. Specifically, we focus on the identification of cellular and molecular mechanisms how reduced calorie intake maintains health, prevents and improves inflammatory diseases, and prolongs healthy life.
Project description:
The focus of my group is to understand the cellular and molecular mechanisms how reduced calorie intake regulates homeostasis and function of the immune system. Recently, we have found that fasting drastically reduces the number of circulating pro-inflammatory monocytes in the blood of humans and mice.1 Interestingly, monocytes accumulated in the bone marrow. Our preliminary data suggest that this is due to the inhibition of monocyte egress from the bone marrow as well as to recruitment from the blood circulation. This phenomenon can be observed not only for monocytes but also for specific other immune cell populations such as naïve B cells and memory T cells.2,3 Intriguingly, memory T cells that have been re-located to the bone marrow display functional modulation and improvement upon egress and re-circulation.2 Hence, our research question is: Are monocytes functionally modified in the bone marrow during fasting? To answer this question, we will establish a monocyte adoptive transfer model in mice and apply next generation sequencing to monocytes in the periphery and the bone marrow. Thus, this project offers the opportunity to the student to develop bench working skills as well as getting familiar with the computational analysis of large transcriptomic datasets.
The aim of the study is to analyze the transcriptional profile of monocytes that returned to the bone marrow from the periphery during fasting.
WP1: Establishing an adoptive monocyte transfer model into fed and fasted mice. It is crucial to the study that monocytes returning to the bone marrow during fasting can be clearly identified. Therefore, we will isolate Ly-6C+ monocytes from the spleen of CD45.1+ mice using the Miltenyi monocyte isolation kit (Fig. 1). Isolated monocytes will be transferred into CD45.2+ mice that will be fed or fasted for 16 hours via intravenous injection. After 4 hours, bone marrow, blood, spleen, and additional organs will be harvested and analyzed for transferred CD45.1+ monocytes using multicolor flow cytometry. Hence, CD45.1/CD45.2 discrimination will enable us to identify and quantify monocytes that are recruited from the blood to the bone marrow. We already obtained the approval of the responsible state office for the animal experiments.
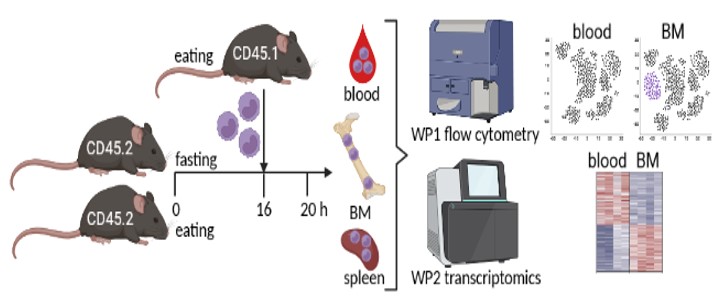
BM = bone marrow.
WP2: Monocyte transcriptomics and computational analysis. In WP2 we will transfer splenic CD45.1+ monocytes into fed and fasted mice as described in WP1 for transcriptional analysis. We will analyze CD45.1+ monocytes that were recruited to the bone marrow, as well as CD45.1+ monocytes that remained in the blood circulation or homed to the spleen. The comparison between monocytes that returned to the bone marrow in fed vs. fasted mice will identify specific fasting-induced modifications. We will also include a monocyte sample from the spleen of a fed mouse for analysis of the pre-transfer state. We will enrich CD45.1+ monocytes from the respective organs using the Miltenyi monocyte isolation kit and subsequently use flow cytometry to sort to high purity for sequencing. Because we do not expect to yield high cell numbers, we will use ultra-low input sequencing approaches and multiplex samples to save resources. The computational analysis will include among others: differentially expressed genes between bone marrow and peripheral monocytes during fasting, gene ontology, KEGG and pathway analysis, upstream regulators of transcriptional changes, prediction of modified cellular functions, analysis of cellular metabolic and transcriptional networks as well as intercellular communication using Ingenuity Pathway analysis and R tools as we have done before.1
In summary, we will investigate functional modifications of monocytes in the bone marrow during fasting using a monocyte transfer model in combination with transcriptional analysis. The results could indicate how fasting reduces the pro-inflammatory potential of monocytes and, thereby, prevents inflammatory disease and prolongs healthy life.
References
- Jordan S, Tung N, Casanova-Acebes M, […], Berres ML, Gallagher EJ, Merad M. Dietary Intake Regulates the Circulating Inflammatory Monocyte Pool. Cell. 2019; 178:1102-1114 e1117. doi: 10.1016/j.cell.2019.07.050.
- Collins, N, Han SJ, Enamorado M, […], McGavern DB, Schwartzberg PL, Belkaid Y. The Bone Marrow Protects and Optimizes Immunological Memory during Dietary Restriction. Cell. 2019; 178:1088-1101 e1015. doi: 10.1016/j.cell.2019.07.049.
- Nagai M, Noguchi R, Takahashi D, […] Takubo K, Dohi T, Hase K. Fasting-Refeeding Impacts Immune Cell Dynamics and Mucosal Immune Responses. Cell. 2019; 178:1072-1087 e1014. doi: 10.1016/j.cell.2019.07.047.

